copick: An open dataset interface and toolkit for collaborative annotation and analysis of cryo-electron tomography data
Apr 17, 2026·,,,,,,,,,·
0 min read
Utz Heinrich Ermel
Jonathan Schwartz
Zhuowen Zhao
Daniel Ji
Ariana Peck
Yue Yu
Mohammadreza Paraan
Bridget Carragher
Achilleas S. Frangakis
Kyle I. S. Harrington
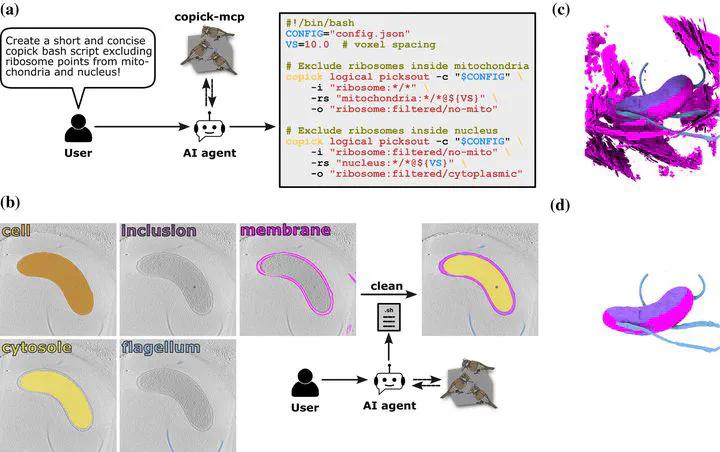 copick-mcp pipeline generation workflow (Figure 4, Ermel et al., 2026, Protein Science)
copick-mcp pipeline generation workflow (Figure 4, Ermel et al., 2026, Protein Science)Abstract
Cryo-electron tomography (cryoET) enables visualization of macromolecular complexes within intact cellular environments. Continued improvements in instrumentation, sample preparation, and data-processing pipelines have increased both the scale and the complexity of cryoET datasets, making manual analysis challenging. To support scalable, collaborative annotation, we developed copick, an open-source dataset application programming interface (API) and accompanying tool suite for cryoET analysis. Copick provides standardized access to tomograms, segmentations, point annotations, meshes, and feature maps across local storage, high-performance computing systems, cloud platforms, and public repositories. Plugins for napari and ChimeraX enable human-in-the-loop workflows for particle picking, segmentation, inspection of machine-learning outputs, and project-level collaboration. A multi-resolution Open Microscopy Environment (OME)-Zarr architecture supports responsive visualization and cross-platform access. Copick additionally provides a Model Context Protocol interface enabling automated generation of annotation-curation pipelines using natural-language instructions. Together, these tools support reproducible, scalable, and collaborative cryoET analysis.
Type
Publication
Protein Science